Monitoring malaria parasite reveals evolving drug resistance and infection history
Scientists have discovered that the malaria parasite Plasmodium vivax is evolving rapidly to adapt to conditions in different geographical locations, in particular to defend itself against widely-used antimalarial drugs. The study, published in Nature Genetics today, provides a foundation for using genomic surveillance to guide effective strategies for malaria control and elimination.
P. vivax is mainly found in Asia and South America, and 2.5 billion people are at risk of infection worldwide. This species of malaria parasite is notoriously difficult to work with and the new study has created one of the largest genomic data sets of this species to date, which is available to all researchers.
The international team of researchers led by Professor Dominic Kwiatkowski from the Wellcome Trust Sanger Institute and Wellcome Trust Centre for Human Genetics, studied the genomes of more than 200 parasite samples from multiple locations across Southeast Asia, identifying the strains carried by each patient and revealing their infection history.
Unlike Plasmodium falciparum, its more widely studied cousin, Plasmodium vivax can remain dormant inside a person's liver for years until it emerges causing a malaria relapse. It is extremely difficult to grow under laboratory conditions, and patients have very low levels of P. vivax DNA in their blood. Only now, with modern DNA sequencing technologies, has it been possible to look in detail at the genetics of the parasite that causes relapsing malaria.
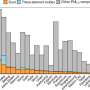
The researchers found that the parasites are evolving rapidly to evade anti-malarial drugs. They revealed that P. vivax evolved differently in Thailand, Cambodia and Indonesia, probably due to the different drug use in each place. This study showed that the genomic signals could help identify local areas of drug resistance as they emerged, information that could support local public health staff.
Dr Richard Pearson, first author of the paper from the Sanger Institute, said: "We can see in the genome that drug resistance is a huge driver for evolution. Intriguingly, in some places, this process appears to be happening in response to drugs used primarily to treat a different malaria parasite, P. falciparum. Although the exact cause isn't known, this is a worrying sign that drug resistance is becoming deeply entrenched in the parasite population."
There are several possible reasons why P. vivax may be evolving to evade drugs used against P. falciparum. Many people carry mixed infections of both species of parasite, so that in treating one species the other automatically gets exposed to the drug. Unsupervised drug use where many people take the most readily available, rather than the most suitable, antimalarial drug may also contribute.
A further key finding was that, when the researchers identified patients who were carrying multiple strains of parasite in their blood, the genomic data made it possible to determine how closely the different strains were related to one another.
Professor Kwiatkowski said: "This means that we can now start to pull apart the genetic complexity of individual Plasmodium vivax infections, and work out whether the parasites came from one or more mosquito bites. It provides a way of addressing fundamental questions about how P. vivax is transmitted and how it persists within a community, and in particular about the biology of relapsing infections."
Professor Ric Price, from the University of Oxford and Menzies School of Health Research, Australia, said: "The front line drug used to treat vivax malaria is chloroquine. Our study shows that the strongest evidence of evolution is in Papua, Indonesia, where resistance of P. vivax to chloroquine is now rampant. These data provide crucial information from which we can start to identify the mechanisms of drug resistance in P. vivax".
Professor Kwiatkowski, added: "For so long, it's not been possible to study P. vivax genomes in detail, on a large-scale, but now we can - and we're seeing the effect that drug use has on how parasites are evolving. In the near future, genomic data will provide more powerful surveillance tools to those who are trying to tackle these problems and make decisions about how to control and eliminate malaria."
More information: Richard Pearson et al. (2016) Genomic analysis of local variation and recent evolution in Plasmodium vivax. Nature Genetics DOI: 10.1038/ng.3599
Related:
Population genomics studies indentify signatures of global dispersal and drug resistance in Plasmodium vivax, Nature Genetics, DOI: 10.1038/ng.3588
Journal information: Nature Genetics
Provided by Wellcome Trust Sanger Institute